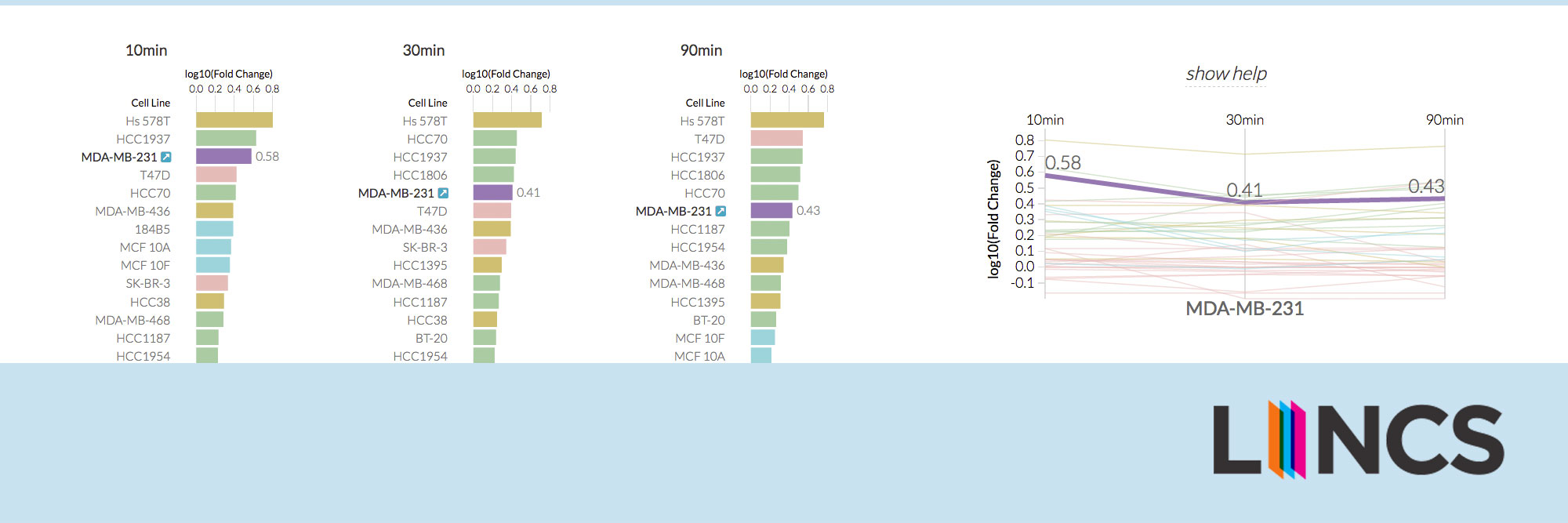
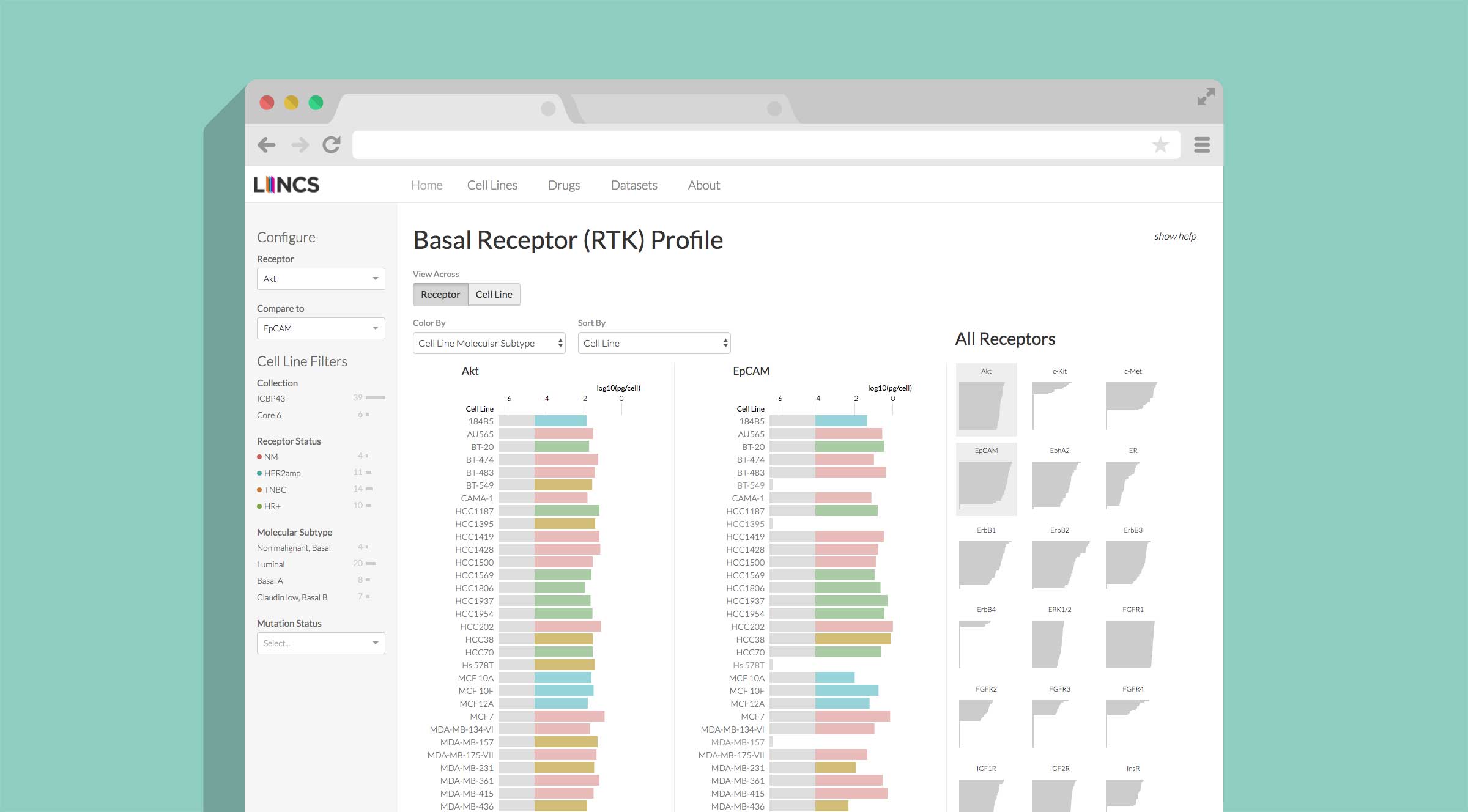
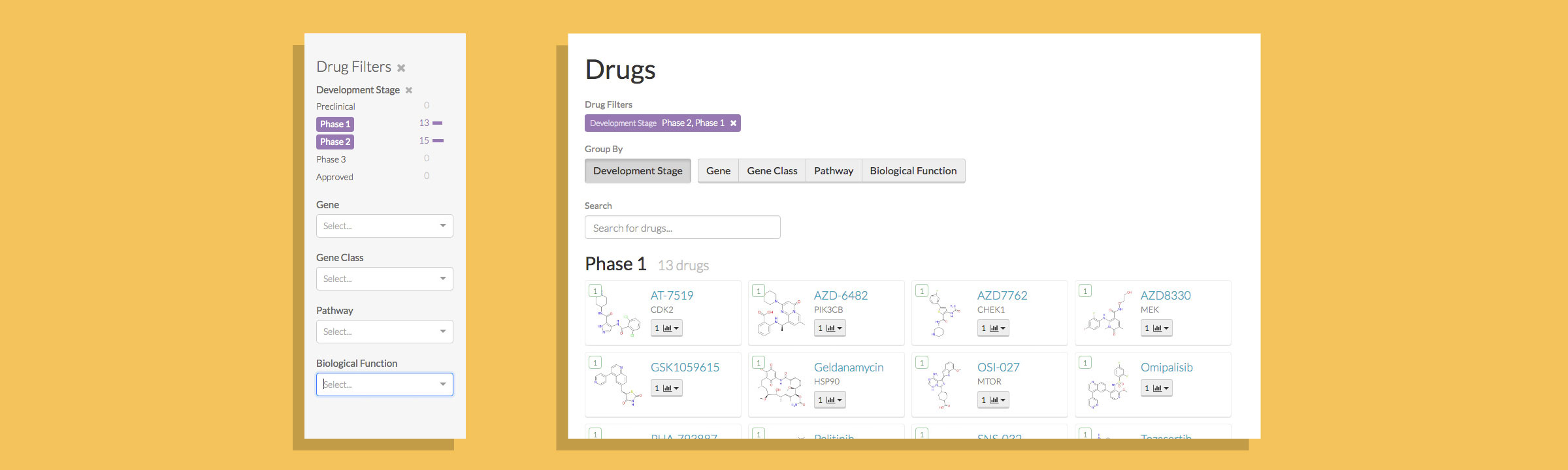
HMS LINCS Breast Cancer Browser
Harvard Medical School
Enabling researchers to investigate breast cancer biology and drug response through exploratory visualizations.
Challenge
Solution
Impact
More work like this at Bocoup
Lyra – Visualization Design Environment
University of Washington Interactive Data Lab
Learn More about Lyra – Visualization Design Environment
JSI Clinic Efficiency Dashboard
JSI Research & Training Institute
Learn More about JSI Clinic Efficiency DashboardContact Us
We'd love to hear from you. Get in touch!